Spot the Differences in Developing Brain Circuits: How Organoids Replay Innate ‘Backbone’ Burst Motifs
If you grew up in Italy with La Settimana Enigmistica, you probably remember those puzzles where two almost-identical drawings hide a handful of tiny differences. If you’re not Italian, the magazine may be different, but the game is the same: you stare, compare… and suddenly you can’t unsee the mismatches.

In a recent Nature Neuroscience paper, van der Molen et al. play a very similar game – but their “drawings” are neural activity patterns recorded from human brain organoids, mouse organoids, neonatal mouse brain slices and 2D cortical cultures. All these preparations are developing networks grown without any sensory input, yet they already display surprisingly structured activity in time. This suggests that some aspects of how circuits fire are not learned from experience, but are preconfigured by the neurodevelopmental program that assembles the network in the first place.
Before playing our scientific “spot the difference” game, we need to know what we’re looking at. The authors recorded extracellular activity with the MaxOne Single-Well High-Density Microelectrode Array (HD-MEA) System, capturing spikes from many units in parallel with high spatial and temporal resolution. That data richness is what lets them zoom in on burst dynamics – moments when the population firing rate shoots up and neurons fire together. And the burst is the signature of strong synchronisation and self-organization: for a few hundred milliseconds, the developing network briefly behaves like a single computational unit.
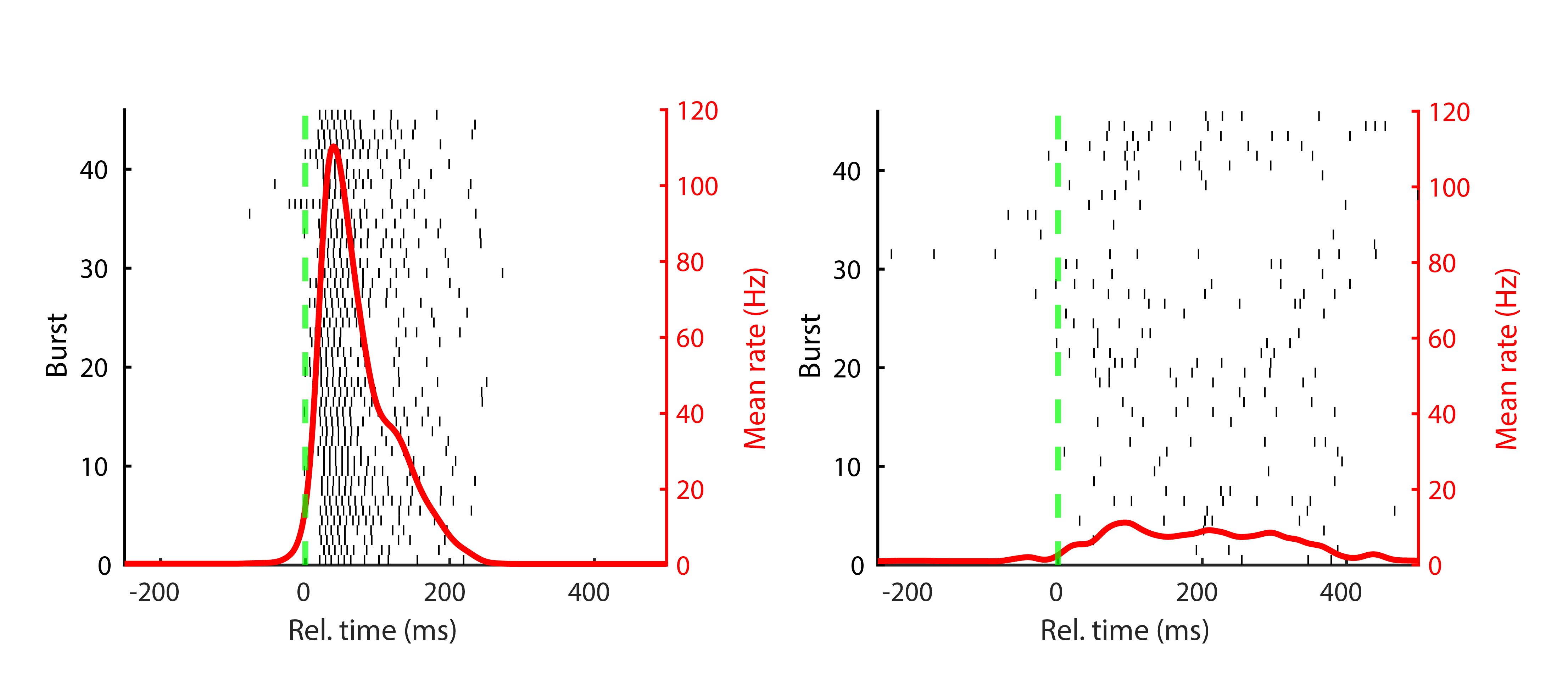
But do all recorded units contribute equally to this orchestrated discharge? That question forms the analytical core of the paper. Each unit was assigned to one of two groups, based on its participation across bursts. A minority of units that consistently help build the burst structure in every detected burst were labelled backbone units; the rest, which join in more variably and less reliably, were grouped as non-rigid units. If there really is an innate firing motif shaped by neurodevelopmental wiring, we expect it to live in this backbone population.
Now, let’s focus on those backbone units and look at their firing pattern across bursts. In the figure below, you see three individual bursts from one of the four preparations; each row is a single backbone unit. How do they look to you – similar, completely different, somewhere in between? And what happens if you average all those backbone burst dynamics together?
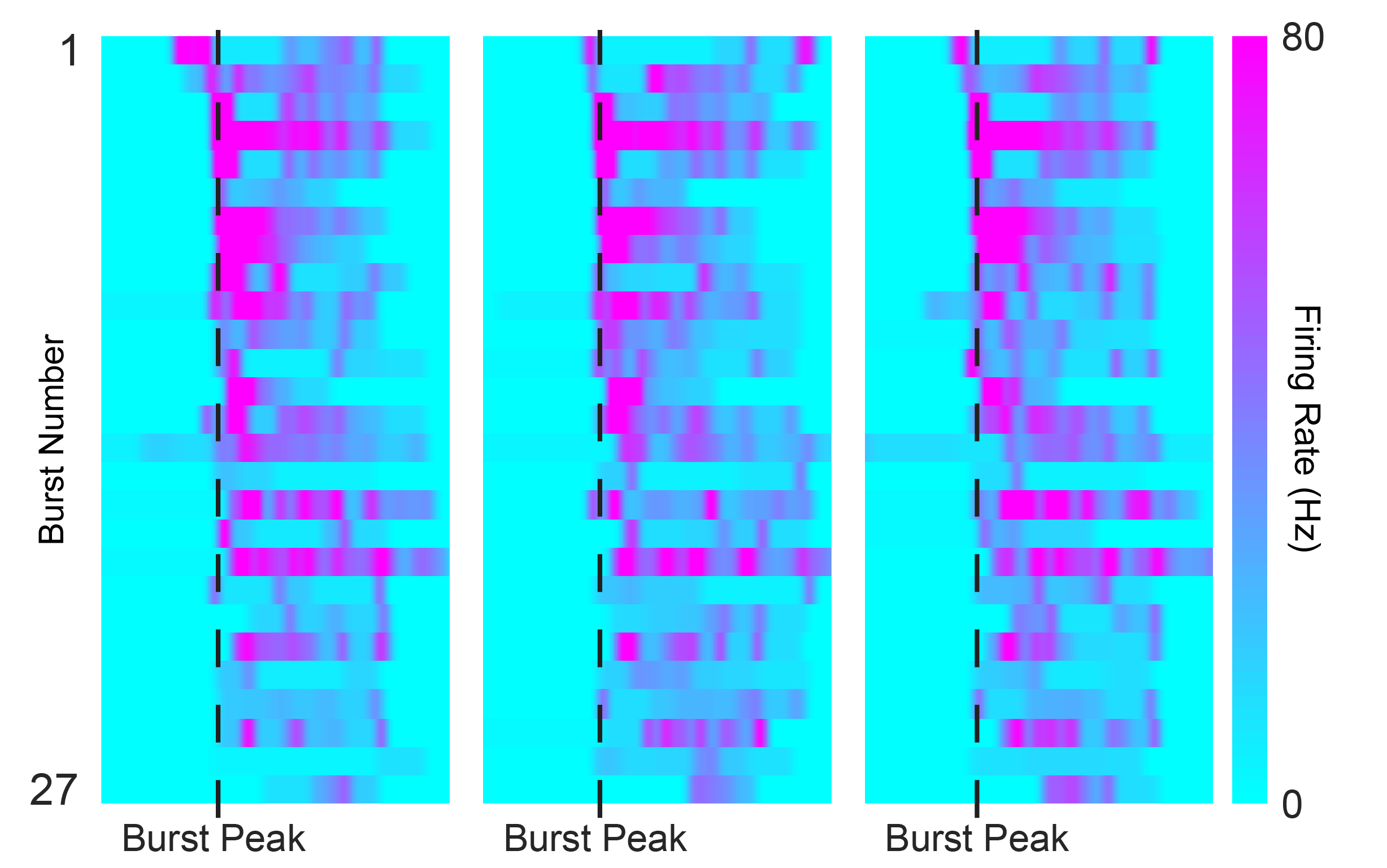
Intuitively, I was expecting the bursts in these immature circuits to be messy and inconsistent, with little burst-to-burst reproducibility. After all, these systems have never been “trained” by sensory experience. The surprise of the paper is that, thanks to the precision and throughput of the MaxWell Biosystems’ HD-MEA recordings, the authors can show that the patterns are far from random: even at early developmental stages, the network keeps replaying a structured temporal pattern.
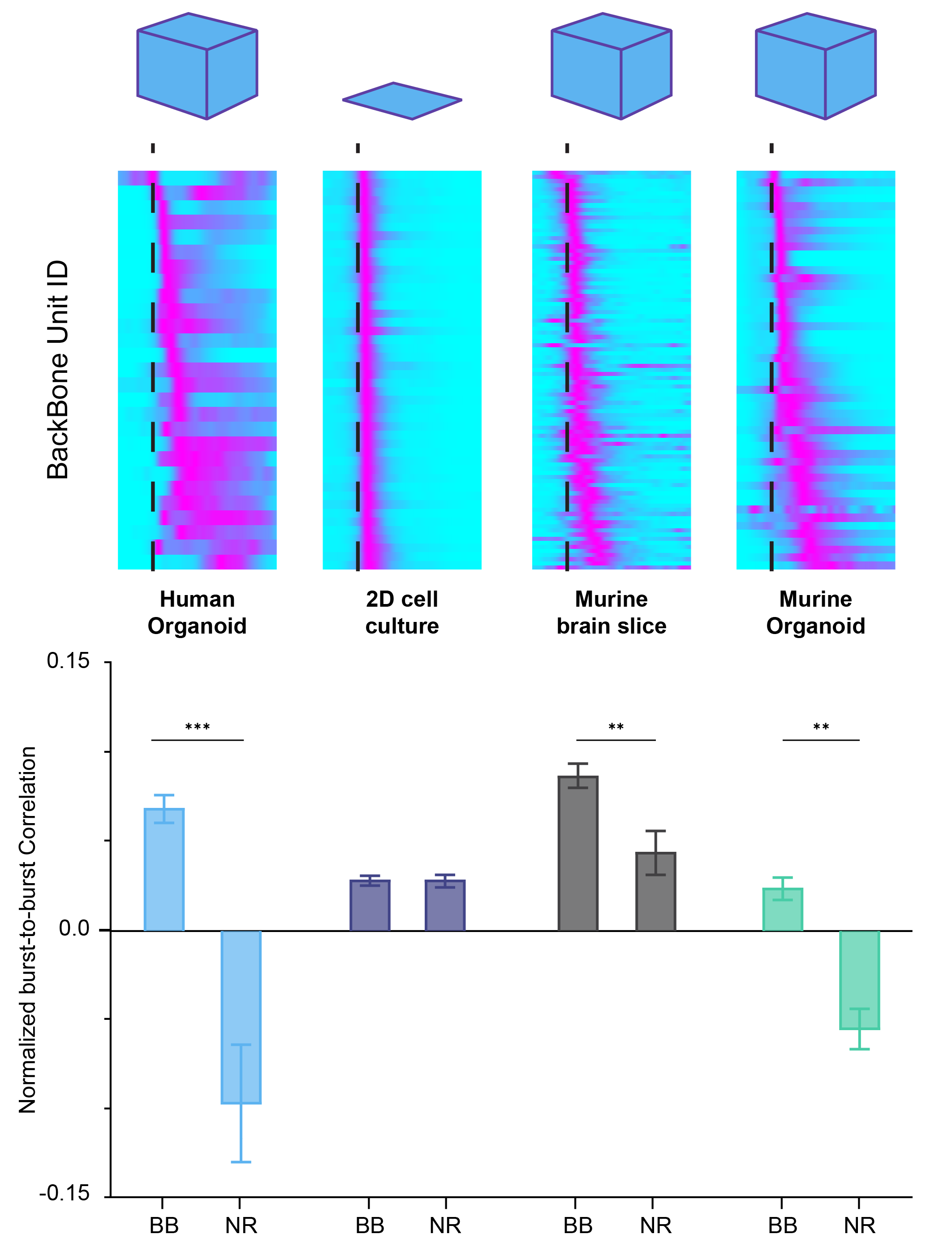
What happens when you average across bursts? We can have a look together in the figure below, each panel shows the average firing dynamics of backbone neurons from the different preparations as a burst unfolds; the dashed green line marks the burst peak. I won’t tell you yet what preparation stand for each panel – that’s your puzzle. Do they all look the same? Ignore that some are more pixelated (that’s just the number of neurons included) and instead follow how the lighter part of the wave shifts in time around the burst peak.
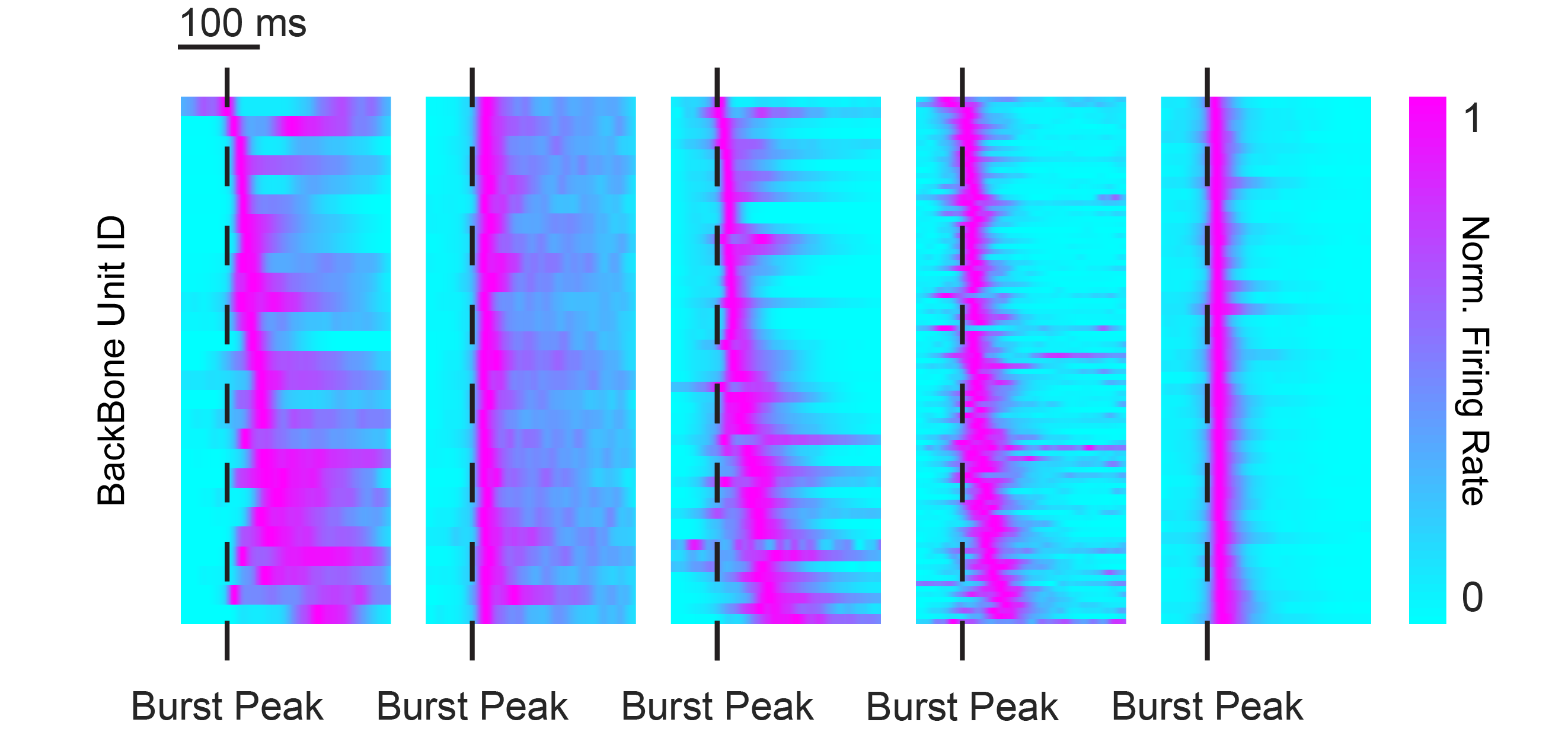
If we try to “pair them up”, some panels look like a very good match, while others clearly don’t.
You may also have noticed that there are five panels, even though we said we had four preparations… and here comes the twist: the second panel is not a real biological preparation, but a shuffled population generated by randomising spikes across units from the first panel. Same overall activity, but the temporal structure is scrambled. This shuffled model, built on top of the same MaxOne HD-MEA dataset, serves as an internal control for each preparation’s burst dynamics and anchors their measure of pattern repeatability.
By running a burst-to-burst cross-correlation analysis for every unit and then normalising that metric against the shuffled version for both backbone and non-rigid units, the authors can elegantly quantify how stable the pattern is across the network – basically, whether bursts are just noisy fireworks or evidence of a fixed, developmentally preconfigured motif in the circuit. By combining these analyses on MaxOne HD-MEA recordings from the four preparations, they can directly show which model truly expresses an innate backbone motif and which does not. That’s what you see in the results figure below.
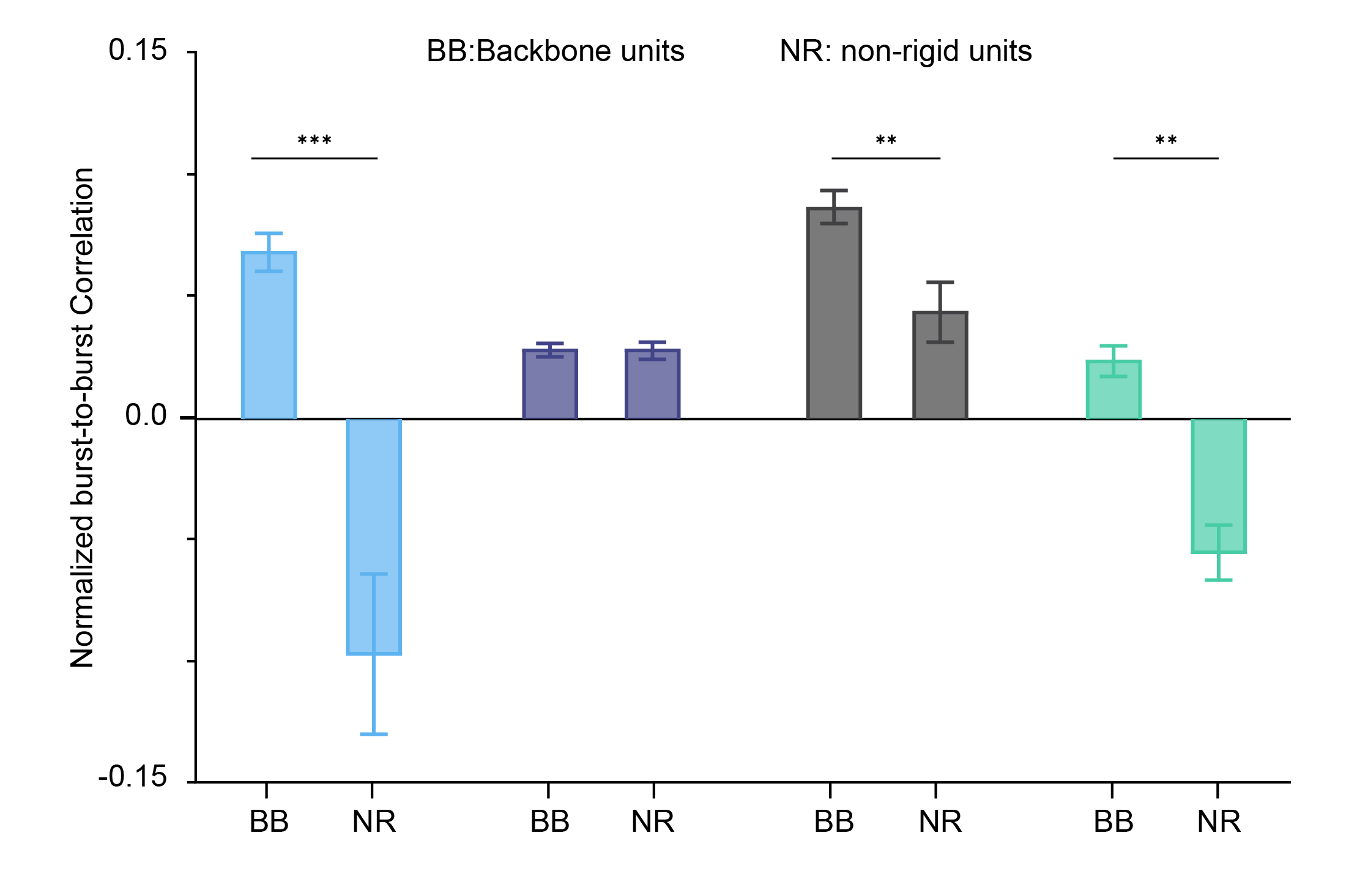
Now I’ll leave you with one last question: who is who? Which of these extracted metrics match the backbone maps you saw before from the MaxOne HD-MEA recording? Which of these preparations recapitulate a fixed, innate backbone motif, and which one fails to do so?
Here we are at the last page of the puzzle magazine – the solutions page, the one you’d normally have to wait a week for, but we’re revealing right now. The model that fails to show a clear neural motif carried by backbone units is the 2D primary neuron culture. This tells us that, unlike organoids and neonatal slices, 2D cultures do not support robust, recurring backbone sequences, making them a poor choice when the goal is to capture recurrent temporal motifs that emerge during cortical circuit development. In contrast, 3D organoids and neonatal slices naturally generate these structured sequences, even in the absence of sensory input, underscoring their value as models of early neuronal circuit assembly.

Dr. Tjitse Van der Molen
The high electrode density of the MaxWell Biosystems' MEAs allows us to resolve the activity of individual neurons across large populations in vitro. In our 3D culture models, this revealed structured firing sequences reminiscent of in vivo circuit dynamics, a finding that became the foundation of this work.
If you’re curious to dive deeper into how these preconfigured sequences were discovered – and how MaxWell Biosystems’ MaxOne Single-Well and MaxTwo Multiwell HD-MEA Systems enable this kind of single-unit, burst-resolved analysis in organoids and other complex developmental preparations – you can explore the original paper and the related work they compare to here Preconfigured neuronal firing sequences in human brain organoids. In fact, they benchmark their findings against data recorded on a MaxTwo Multiwell HD-MEA System in an independent lab, and still see the same story: strong backbone motifs in 3D developmental models, and none in 2D cultures. That convergence across preparations, developmental stages, laboratories and MaxWell Biosystems’ platforms is a neat built-in validation of the reliability and reproducibility of the technology. The next time you play spot the differences, remember that some of the most interesting ones live not in pixels, but in milliseconds of activity that the right chip can reveal.
Related
Resources
MaxOne
Versatility and functionality in one compact device

MaxTwo
Maximize your cell functional assays

MaxLab Live
All-in-one Software
Our Technology
Multi-/Micro-electrode Arrays

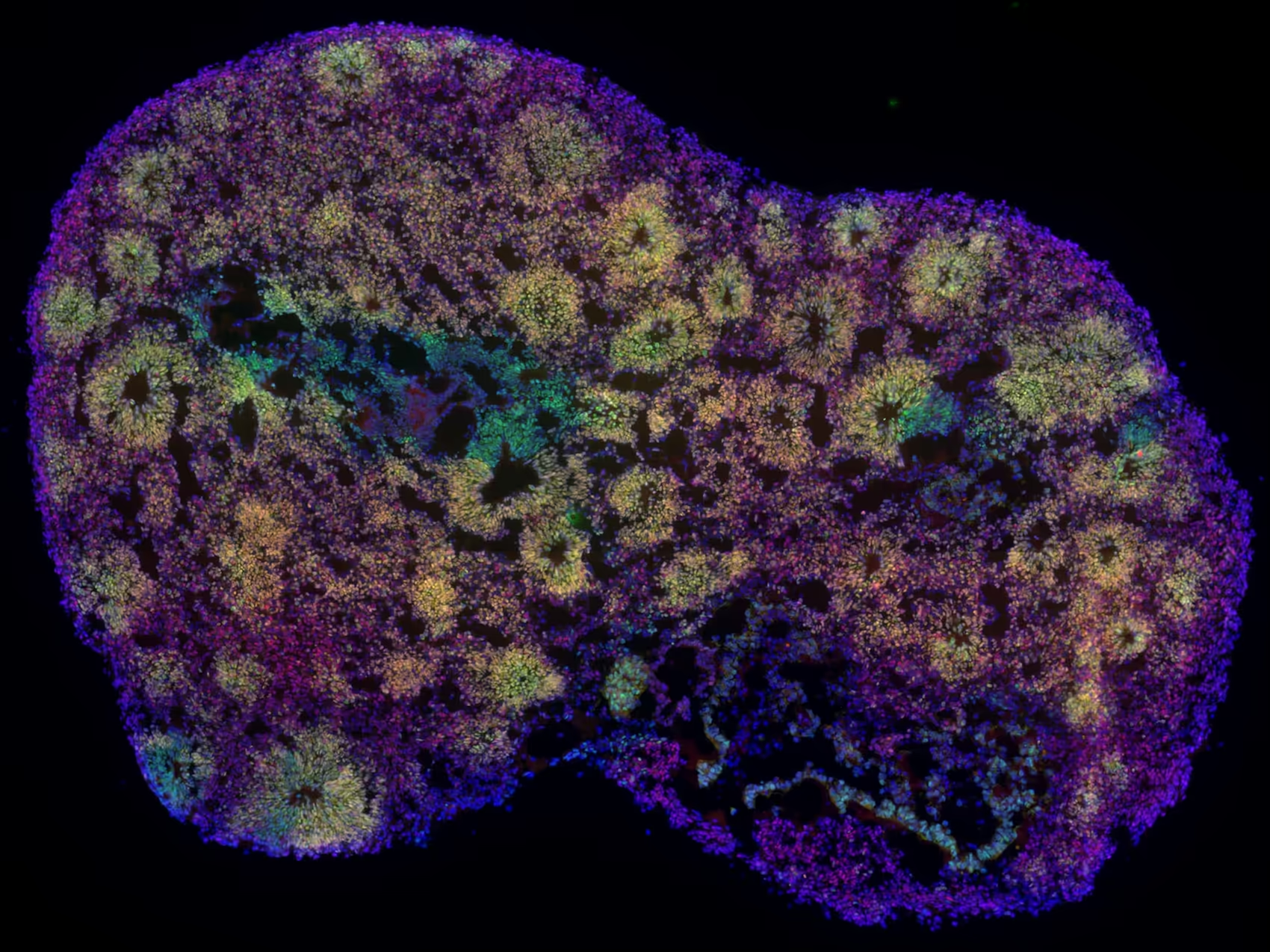
Organoids
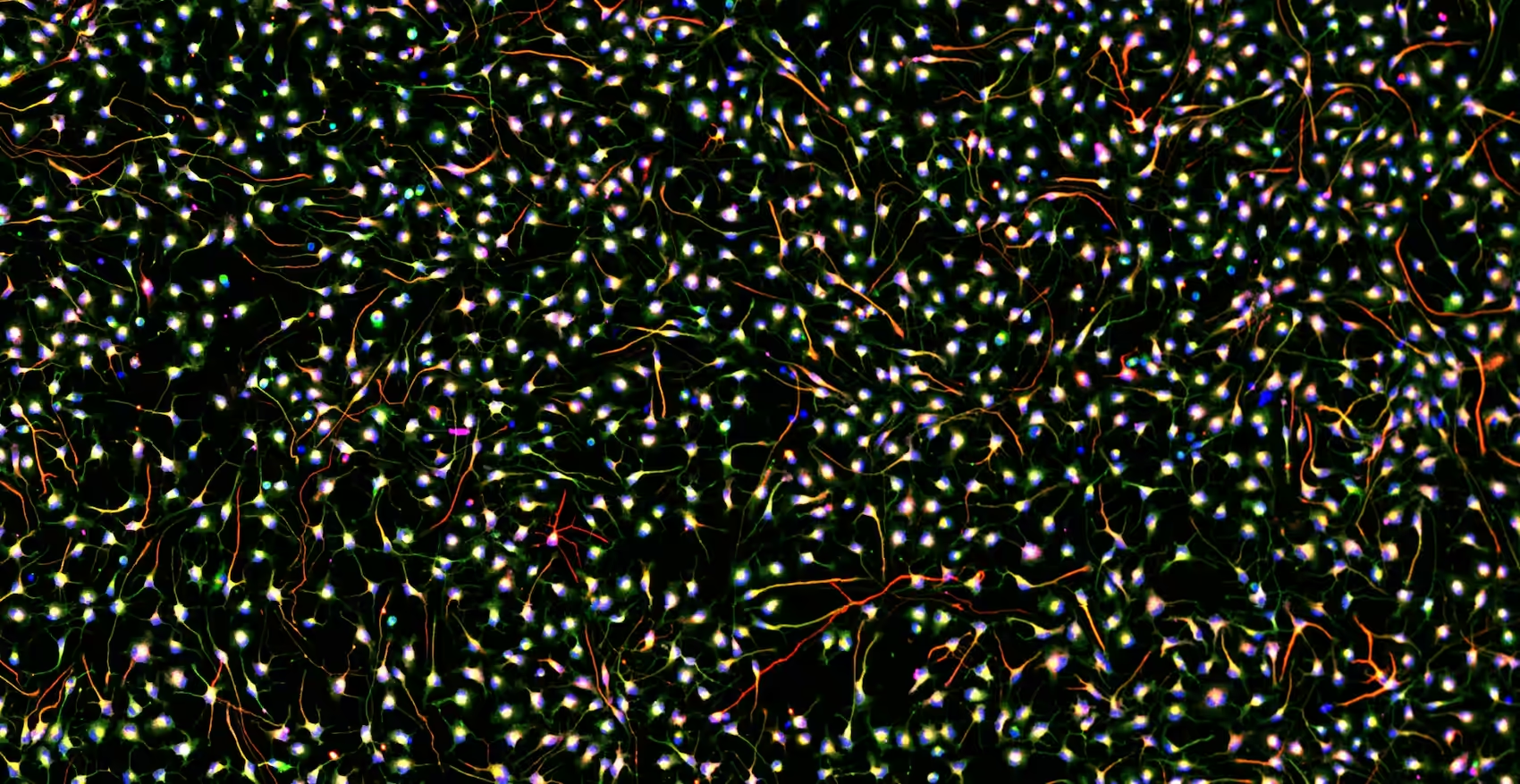
Neuronal Cell Cultures
Functional Phenotyping